Project highlights
- Investigate coral adaptation to the world’s warmest reefs. In this project, you will be analysing gene copy number variation, an often overlooked but important source of genomic variation.
- You will develop cutting-edge bioinformatic skills analysing short- and long-read sequencing datasets.
- You will be working at the forefront of ecological genomics research whilst contributing to a growing body of research working to conserve coral reefs for future generations.
Overview
Coral reefs have suffered dramatic declines over the past three decades due to a myriad of natural and anthropogenic stressors. As the threats to coral reefs are expected to increase under climate change, it is imperative that we understand the ecological and evolutionary mechanisms underpinning coral stress tolerance. An improved understanding of corals’ adaptive capacity will help improve forecasts of reef futures in response to climate change, identify vulnerable populations for conservation, and enhance reef restoration efforts.
To date, research on the genomic basis for coral adaptation has focussed on using single nucleotide polymorphisms (SNPs) to identify regions of the genome under selection (e.g., Smith et al., 2022). The focus on SNPs has largely resulted from the ease at which they can be obtained through short-read sequencing technologies, however, SNPs are only one of many types of genomic polymorphisms (e.g., inversions, translocations, copy number variation). Copy number variation (CNV) is a form of genomic variation that arises from the duplication of genomic segments, which can alter gene expression through dosage effects, as well as provide substrate for adaptive evolution. CNV has been recognised as an important source of variation between coral species (Noel et al., 2023), yet remains understudied at the species level (i.e., intraspecific variation). This project will take advantage of recent technological and methodological advances to investigate whether copy number variation (CNV) is an important driver of coral adaptation to distinct thermal environments.
In this project, you will quantify the extent of CNV among individuals of the same coral species collected from the extreme environmental gradient located between the Persian/Arabian Gulf and the Gulf of Oman. You will identify gene family expansions associated thermal and salinity gradients, and assess how CNV at the genomic level impacts the transcriptome/proteome.
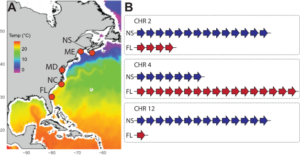
Figure 1: Our recent research on the starlet sea anemone (Nematostella vectensis) along the Atlantic Coast of the U.S. has revealed extensive copy number variation (CNV) between populations (e.g., Smith et al. 2023). In this project, you will be using similar techniques to investigate CNV differences in corals.
Host
University of WarwickTheme
- Organisms and Ecosystems
Supervisors
Project investigator
Dr Ed Smith (University of Warwick, [email protected])
Co-investigators
Prof Robin Allaby (University of Warwick, [email protected])
How to apply
- Each host has a slightly different application process.
Find out how to apply for this studentship. - All applications must include the CENTA application form. Choose your application route
Methodology
Copy number variation among individuals of the same coral species will be compared using whole genome sequencing. There is freedom for the student to target particular genes of interest and quantify copy number differences and sequence divergence using multiple approaches including qPCR, amplicon sequencing, and comparative genomics (see Smith et al. 2023). In order to gain functional insights, we plan to supplement these analyses with additional approaches that could include common-garden experiments, transcriptomics, proteomics, and/or microscopy.
Training and skills
Students will be awarded CENTA2 Training Credits (CTCs) for participation in CENTA2-provided and ‘free choice’ external training. One CTC equates to 1⁄2 day session and students must accrue 100 CTCs across the three years of their PhD.
Training will be provided in a range of different techniques, as required, including molecular biology (e.g., high molecular weight DNA extractions, qPCR, sequencing library preparation), bioinformatics (e.g., genome assembly, comparative genomics), and experimental design.
Partners and collaboration
This project will involve collaboration with Dr. John Burt and members of Burt Lab at New York University Abu Dhabi.
Further details
Further details on how to contact the supervisor for this project and how to apply for this project can be found here:
Please see the lab page here.
To learn more about the project, contact Ed Smith ([email protected]). Please include a CV, details of past research, and outline your interest in the project.
To apply to this project:
- You must include a CENTA studentship application form, downloadable from: CENTA Studentship Application Form 2024.
- You must include a CV with the names of at least two referees (preferably three) who can comment on your academic abilities.
- Please submit your application and complete the host institution application process via: https://warwick.ac.uk/fac/sci/lifesci/study/pgr/studentships/nerccenta/ Complete the online application form – selecting course code P-C1PB (Life Sciences PhD); from here you will be taken through to another screen where you can select your desired project. Please enter “NERC studentship” in the Finance section and add Nikki Glover, [email protected] as the scholarship contact. Please also complete the CENTA application form 2024 and submit via email to [email protected]. Please quote CENTA 2024-W3 when completing the application form.
Applications must be submitted by 23:59 GMT on Wednesday 10th January 2024.
Possible timeline
Year 1
Genome-wide characterisation of intraspecific CNV using population genomics WGS data or genome assemblies.
Year 2
In-depth analyses of target genes across different environments.
Year 3
Integrating experimental approaches to gain mechanistic insights into the functional effects of genomic copy number variation.
Further reading
Dorant, Y., Cayuela, H., Wellband, K., Laporte, M., Rougemont, Q., Mérot, C., Normandeau, E., Rochette, R. and Bernatchez, L. (2020) Copy number variants outperform SNPs to reveal genotype–temperature association in a marine species. Molecular Ecology, 29(24), pp.4765-4782.
Noel, B., Denoeud, F., Rouan, A., Buitrago-López, C., Capasso, L., Poulain, J., Boissin, E., Pousse, M., Da Silva, C., Couloux, A. and Armstrong, E. (2023) Pervasive tandem duplications and convergent evolution shape coral genomes. Genome Biology, 24(1), pp.1-38.
Smith, E.G., Hazzouri, K.M., Choi, J.Y., Delaney, P., Al-Kharafi, M., Howells, E.J., Aranda, M. and Burt, J.A. (2022) Signatures of selection underpinning rapid coral adaptation to the world’s warmest reefs. Science advances, 8(2), p.eabl7287.
Smith, E.G., Surm, J.M., Macrander, J., Simhi, A., Amir, G., Sachkova, M.Y., Lewandowska, M., Reitzel, A.M. and Moran, Y. (2023) Micro and macroevolution of sea anemone venom phenotype. Nature Communications, 14(1), p.249.